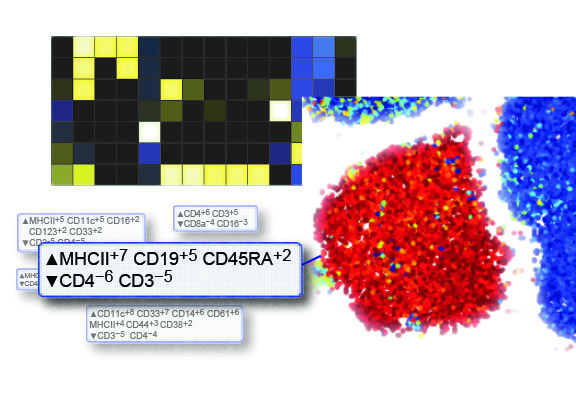
Marker Enrichment Modeling (MEM) is an open-source R package developed by Kirsten E. Diggins and Jonathan M. Irish, Ph.D., at Vanderbilt University’s Meiler Lab to quantify marker enrichment in cell populations. Designed for computational biologists, MEM characterizes cell subsets in high-dimensional data, such as mass cytometry or single-cell RNA sequencing (scRNA-seq), enabling researchers to identify unique features and advance discoveries in immunology and cancer.
Key features include:
- Multivariate modeling to quantify feature enrichment without predefined thresholds.
- Compatibility with R environments on Windows, MacOS, and UNIX systems.
- Inclusion of source code, metadata, vignette, and sample script for flexible analysis.
Benefits: MEM outperforms traditional differential expression and clustering methods by highlighting what makes cell subsets unique, streamlining analysis and supporting publication-ready insights. Its open-source design accelerates research in immune response characterization and cancer studies, making it a critical tool for academic and commercial applications.
Request the free Academic License or a Commercial License Quote on our licensing page. For more details, see our FAQs or contact us with your question using our Contact Us page.
